Can Google DeepMind unlock the mystery of life’s origin?
Vicente Soriano
UNIR Health Sciences School and Medical Center, Madrid, Spain
*Correspondence: Vicente Soriano. Email: vicente.soriano@unir.net
Received: 28-01-2026
Accepted: 03-02-2026
DOI: 10.24875/AIDSRev.M26000093
Available online: 04-02-2026
AIDS Rev. 2026;28(1):42-43
Contents
The central dogma of molecular biology was formulated by Francis Crick in 1958. Life is sustained by a unidirectional flow of genetic information within each cell: DNA replicates, is transcribed into messenger RNA, and this is translated into amino acids that make up proteins (Fig. 1).
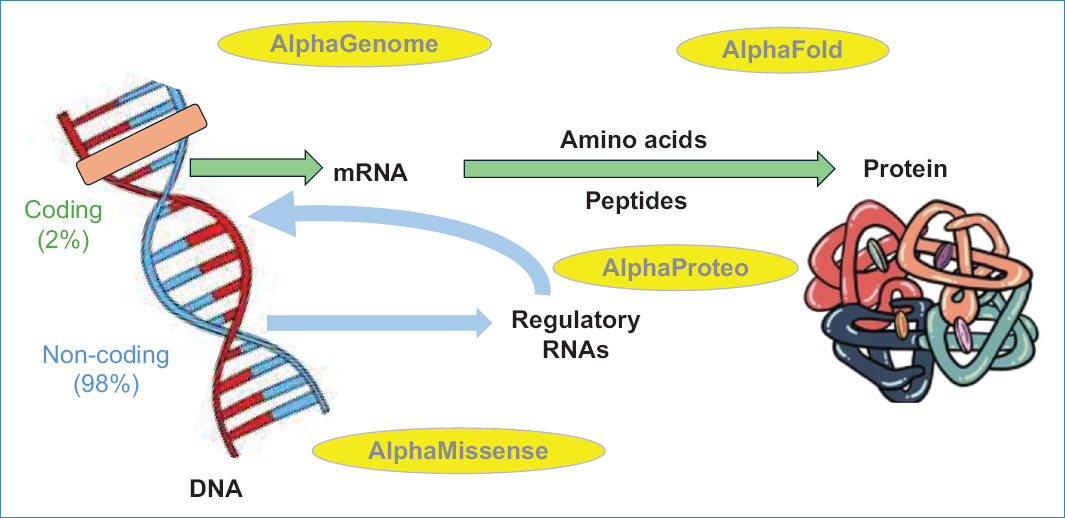
Figure 1. Central dogma of molecular biology and Google DeepMind software tools.
In 2010, DeepMind was founded in the United Kingdom with the aim of developing advanced computing and artificial intelligence (AI) applications. In 2014, it was acquired by Google. At present, Google DeepMind is an AI research biotech company within the Alphabet (Google) group.
Today, in computational biology, Google DeepMind’s main projects focus on understanding the mechanisms of action and function of the elements of the central dogma. These programs decipher life’s mechanisms and, as a result, could potentially enable us to intervene and modify them. With this knowledge, preventing diseases and developing new treatments may become possible.
The “Four Alphas” of Google DeepMind
During the last 5 years, Google DeepMind has developed four major biotechnology tools: AlphaFold, AlphaMissense, AlphaGenome, and AlphaProteo.
AlphaFold was introduced in 2018, but its more accurate version is from 2020 (Jumper et al., Nature 2021). It has revolutionized the prediction of protein three-dimensional structures from amino acid sequences. Its designers were recognized with the 2024 Nobel Prize in Chemistry.
With AlphaMissense, the effects of “missense” mutations – those that cause amino acid changes – can be predicted, revealing whether a genetic variant may be harmful or benign. This rapid prediction aids genetic diagnosis, such as identifying variants behind rare diseases (Cheng et al., Science 2023).
AlphaGenome, the most recent AI program, analyses long DNA sequences (up to 1 million bases) to predict the effects of mutations on gene regulation. By doing so, it advances understanding of physiological traits, genetic diseases, developmental biology (embryology), and personalized medicine (Avsec et al., Nature 2026).
While AlphaMissense targets coding DNA (2% of the genome), AlphaGenome focuses on the remaining 98%, which is regulatory in nature (Lozano-Villalba and Puthanveetti, J Biol Chem 2026) (Fig. 2).
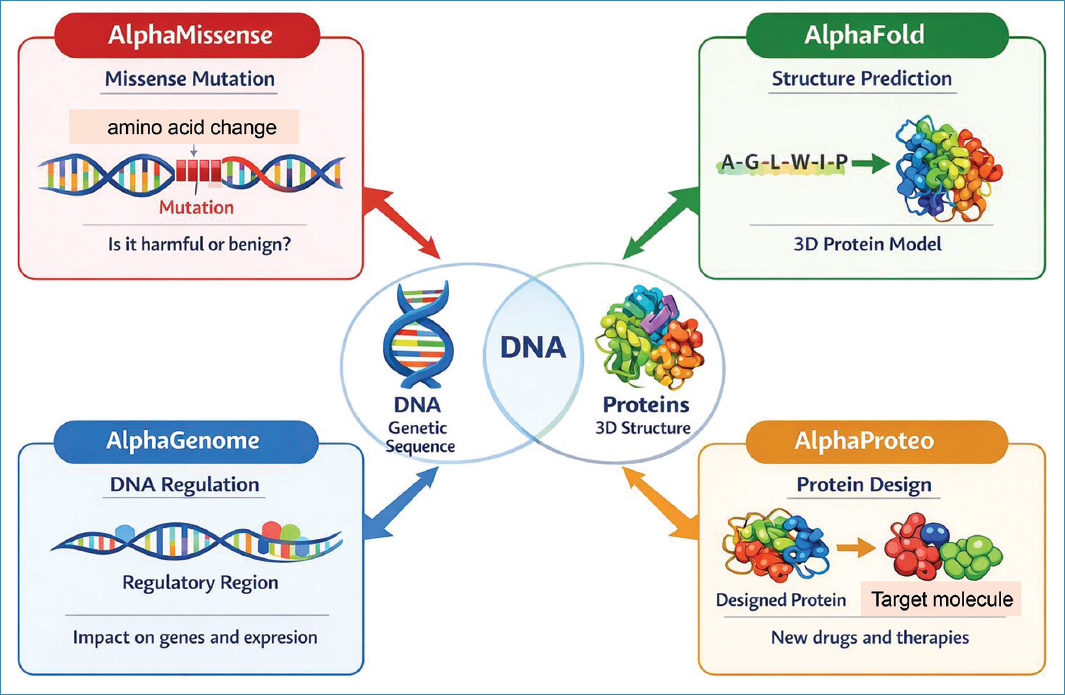
Figure 2. Google DeepMind software tools for deciphering and prediction of molecular biology pathways.
AlphaProteo is an AI program that helps design entirely new proteins, tailored to bind target molecules – acting, for example, as virus cell receptor antagonists or enzyme inhibitors. This powerful tool can accelerate drug discovery (Zambaldi et al., arXiv 2024).
With these advancements, no doubt come pressing ethical questions: what could it mean to create proteins never before seen in nature? How can we safeguard this technology so it is not misused to create artificial pathogens?
Lights and shadows of the new AI biotechnology tools
Advanced deep learning techniques and neural networks power each of these biotech software programs, enabling them to tackle molecular biology challenges infeasible for standard experiments or complex computations. Thanks to the availability of these biotech tools, scientific breakthroughs and the acceleration of biomedical knowledge are now possible.
So far, Google DeepMind has not made public the code or the variable weights used in its bioinformatics models. Ultimately, only open and complete access to these new biotech tools can provide a better understanding and safe biomedical applications (García-González and Golewski, Trends Genet 2026).